Estimate single layer MGM network with bootstrap centrality, bridge metrics, clustering, and (optionally) community score loadings
Source:R/mixMN.R
mixMN.RdEstimates a single layer Mixed Graphical Model (MGM) network on the original data, using the estimation framework implemented in the mgm package, and performs non-parametric bootstrap (row resampling) to compute centrality indices, bridge metrics, clustering stability, and quantile regions for node metrics and edge weights. Optionally, the function computes community score loadings (for later prediction on new data) and can bootstrap the corresponding loadings.
Usage
mixMN(
data,
reps = 100,
scale = TRUE,
lambdaSel = c("CV", "EBIC"),
lambdaFolds = 5,
lambdaGam = 0.25,
alphaSeq = 1,
alphaSel = "CV",
alphaFolds = 5,
alphaGam = 0.25,
k = 2,
ruleReg = "AND",
threshold = "LW",
overparameterize = FALSE,
thresholdCat = TRUE,
quantile_level = 0.95,
covariates = NULL,
exclude_from_cluster = NULL,
treat_singletons_as_excluded = FALSE,
seed_model = NULL,
seed_boot = NULL,
cluster_method = c("louvain", "edge_betweenness", "fast_greedy", "infomap",
"label_prop", "leading_eigen", "leiden", "optimal", "spinglass", "walktrap"),
cluster_args = list(),
compute_loadings = TRUE,
boot_what = c("general_index", "bridge_index", "excluded_index", "community",
"loadings"),
save_data = FALSE,
progress = TRUE
)Arguments
- data
A
data.frame(n x p) with variables in columns. Variables may be numeric, integer, logical, or factors. Character and Date/POSIXt variables are not supported and must be converted prior to model fitting. Variable types are internally mapped to MGM types as follows: continuous numeric (double) variables are treated as Gaussian; integer variables are treated as Poisson unless they take only values in {0,1}, in which case they are treated as binary categorical; factors and logical variables are treated as categorical. Binary categorical variables (two-level factors and logical variables) are internally recoded to {0,1} for model fitting. The original input data are not modified.- reps
Integer (>= 0). Number of bootstrap replications.
- scale
Logical; if
TRUE(default) Gaussian variables (type == "g") are z-standardized internally bymgm(). Usescale = FALSEif your data are already standardized.- lambdaSel
Method for lambda selection:
"CV"or"EBIC".- lambdaFolds
Number of folds for CV (if
lambdaSel = "CV").- lambdaGam
EBIC gamma parameter (if
lambdaSel = "EBIC").- alphaSeq
Alpha parameters of the elastic net penalty (values between 0 and 1).
- alphaSel
Method for selecting the alpha parameter:
"CV"or"EBIC".- alphaFolds
Number of folds for CV (if
alphaSel = "CV").- alphaGam
EBIC gamma parameter (if
alphaSel = "EBIC").- k
Integer (>= 1). Order of modeled interactions.
- ruleReg
Rule to combine neighborhood estimates:
"AND"or"OR".- threshold
Threshold below which edge-weights are set to zero: Available options are
"LW","HW", or"none"."LW"applies the threshold proposed by Loh & Wainwright;"HW"applies the threshold proposed by Haslbeck & Waldorp;"none"disables thresholding. Defaults to"LW".- overparameterize
Logical; controls how categorical interactions are parameterized in the neighborhood regressions. If
TRUE, categorical interactions are represented using a fully over-parameterized design matrix (i.e., all category combinations are explicitly modeled). IfFALSE, the standardglmnetparameterization is used, where one category serves as reference. For continuous variables, both parameterizations are equivalent. The default isFALSE. The over-parameterized option may be advantageous when distinguishing pairwise from higher-order interactions.- thresholdCat
Logical; if
FALSEthresholds of categorical variables are set to zero.- quantile_level
Level of the central bootstrap quantile region (default
0.95). Must be a single number between 0 and 1.- covariates
Character vector. Variables used as adjustment covariates in model estimation.
- exclude_from_cluster
Character vector. Nodes excluded from community detection (in addition to
covariates).- treat_singletons_as_excluded
Logical; if
TRUE, singleton communities (size 1) are treated as excluded nodes when computing bridge metrics.- seed_model
Optional integer seed for reproducibility of the initial MGM fit.
- seed_boot
Optional integer seed passed to
future.applyfor reproducibility of bootstrap replications.- cluster_method
Community detection method used on the clustering graph. Either a character string naming one of the built-in methods
"louvain","edge_betweenness","fast_greedy","infomap","label_prop","leading_eigen","leiden","optimal","spinglass","walktrap", or a user-supplied function.If a function is supplied, it must accept a graph through argument
graphand return either an igraphcommunitiesobject, a list with componentmembership, or a membership vector of length equal to the number of nodes used for clustering.- cluster_args
Named list of additional arguments passed to the selected community detection method. For example,
stepsfor"walktrap",nb.trialsfor"infomap",spinsfor"spinglass". Ignored if not relevant for the selected method.- compute_loadings
Logical; if
TRUE(default), compute community loadings (EGAnet::net.loads). Only supported for Gaussian, Poisson, and binary categorical nodes; otherwise loadings are skipped and the reason is stored incommunity_loadings$reason.- boot_what
Character vector specifying which quantities to bootstrap. Valid options are:
"general_index"(centrality indices),"bridge_index"(bridge metrics for nodes in communities),"excluded_index"(bridge metrics for nodes treated as excluded),"community"(community memberships),"loadings"(community loadings, only ifcompute_loadings = TRUE), and"none"(skip all node-level bootstrap: only edge-weight bootstrap is performed ifreps > 0).- save_data
Logical; if
TRUE, store the original data in the output object.- progress
Logical; if
TRUE(default), show a bootstrap progress bar.
Value
An object of class "mixMN_fit", that is a list with
the following top-level components:
callThe matched function call.
settingsList of main settings used in the call, including
reps,cluster_method,cluster_args,covariates,exclude_from_cluster,treat_singletons_as_excluded, andboot_what.data_infoList with information derived from the input data used for model setup:
mgm_type_level(data frame with one row per variable, reporting the original R class and the inferred MGMtypeandlevel, as used in the call tomgm::mgm), andbinary_recode_map(named list describing the mapping from original binary labels to the internal {0,1} coding used for model fitting).modelList with:
mgm(the fittedmgmobject),nodes(character vector of all node names),n(number of observations),p(number of variables), anddata(ifsave_data = TRUE).graphList describing the graph:
igraph(an igraph object built onkeep_nodes_graph, with edge attributesweight,abs_weight,signand vertex attributemembershipfor communities),keep_nodes_graph(nodes retained in the graph and all node-level metrics), andkeep_nodes_cluster(nodes used for community detection).communitiesList describing community structure with:
original_membership(integer vector of community labels onkeep_nodes_cluster),groups(factor of community labels actually used for bridge metrics, optionally with singletons treated as excluded),palette(named vector of colors per community), andboot_memberships(list of bootstrap memberships if"community"is requested inboot_what, otherwise an empty list).statisticsList with node- and edge-level summaries:
nodeis a list with:true(data frame with one row per node inkeep_nodes_graph, containing the node name and metricsstrength,ei1,closeness,betweenness,bridge_strength,bridge_betweenness,bridge_closeness,bridge_ei1,bridge_ei2, and for nodes treated as excluded from communities alsobridge_strength_excluded,bridge_betweenness_excluded,bridge_closeness_excluded,bridge_ei1_excluded,bridge_ei2_excluded);boot(list of bootstrap matrices for each metric, each of dimensionreps x length(keep_nodes_graph), possiblyNULLif the metric was not requested or ifreps = 0); andquantile_region(list of quantile regions for each node metric, onep x 2matrix per metric, with columns corresponding to the lower and upper quantile bounds implied byquantile_level, orNULLif no bootstrap was performed).edgeis a list with:true(data frame with columnsedgeandweightfor all unique undirected edges amongkeep_nodes_graph);boot(matrix of bootstrap edge weights of dimensionn_edges x reps); andquantile_region(matrix of quantile regions for edge weights,n_edges x 2, with columns corresponding to the lower and upper bootstrap quantile bounds, orNULLifreps = 0).community_loadingsList containing community-loading information (based on
EGAnet::net.loads) for later community-score computation on new data:nodes(nodes used for loadings),wc(integer community labels aligned withnodes),true(matrix of standardized loadings, nodes x communities, orNULLif loadings were not computed.),boot(list of bootstrap loading matrices, one per replication, orNULLif not bootstrapped),available(logical indicating whether loadings were computed),reason(character string explaining why loadings were not computed, orNULLifavailable = TRUE),non_scorable_nodes(character vector of nodes in the community subgraph that prevented loadings from being computed (e.g., categorical variables with >2 levels), otherwise empty).
Details
This function does not call future::plan(). To enable
parallel bootstrap, set a plan (e.g. future::plan(multisession))
before calling mixMN(). If boot_what is "none" and
reps > 0, node-level metrics are not bootstrapped but edge-weight
bootstrap and corresponding quantile regions are still computed.
References
Haslbeck, J. M. B., & Waldorp, L. J. (2020). mgm: Estimating Time-Varying Mixed Graphical Models in High-Dimensional Data. Journal of Statistical Software, 93(8). doi:10.18637/jss.v093.i08
Loh, P. L., & Wainwright, M. J. (2012). Structure estimation for discrete graphicalmodels: Generalized covariance matrices and their inverses. NIPS
Examples
data(bacteremia)
df <- bacteremia[, !names(bacteremia) %in% "BloodCulture"]
fit <- mixMN(
data = df,
lambdaSel = "EBIC",
lambdaGam = 0.25,
reps = 0,
seed_model = 42,
cluster_method = "louvain",
covariates = c("AGE", "SEX"),
compute_loadings = FALSE,
progress = FALSE
)
fit
#> MixMashNet fit
#> Type: Single layer MGM (mixMN)
#> Data: 7420 subjects x 15 variables
#> Graph: 13 nodes, 38 edges
#> Communities: 4
#> Covariates (adjusted for): AGE, SEX
#> Lambda selection: EBIC
#> Community detection: louvain
#> Bootstrap replications: 0
#> Bootstrapped quantities: none (reps = 0)
#> Data info:
#> - Inferred as 'c' (categorical): SEX
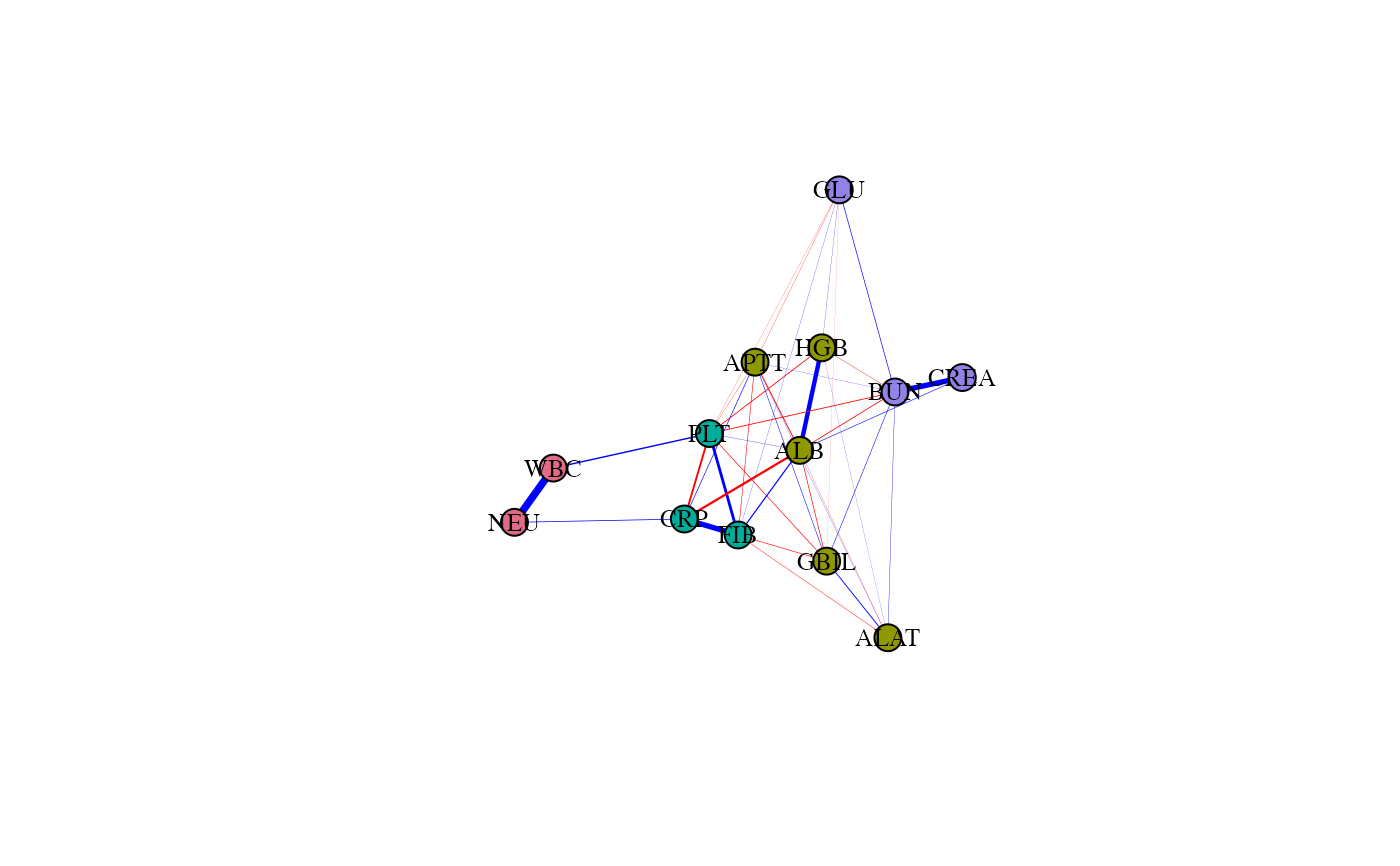
# Plot the estimated network
set.seed(1)
plot(fit)
 # \donttest{
fit_b <- mixMN(
data = df,
lambdaSel = "EBIC",
lambdaGam = 0.25,
reps = 5,
seed_model = 42,
seed_boot =42,
cluster_method = "louvain",
covariates = c("AGE", "SEX"),
boot_what = "community",
compute_loadings = FALSE,
progress = FALSE
)
#>
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
#> Warning: package ‘future’ was built under R version 4.5.2
#> Total computation time: 5.1 seconds (~ 0.08 minutes).
# Plot the membership stability
plot(fit_b, what = "stability", cutoff = 0.7)
# \donttest{
fit_b <- mixMN(
data = df,
lambdaSel = "EBIC",
lambdaGam = 0.25,
reps = 5,
seed_model = 42,
seed_boot =42,
cluster_method = "louvain",
covariates = c("AGE", "SEX"),
boot_what = "community",
compute_loadings = FALSE,
progress = FALSE
)
#>
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
|
| | 0%
|
|----- | 7%
|
|--------- | 13%
|
|-------------- | 20%
|
|------------------- | 27%
|
|----------------------- | 33%
|
|---------------------------- | 40%
|
|--------------------------------- | 47%
|
|------------------------------------- | 53%
|
|------------------------------------------ | 60%
|
|----------------------------------------------- | 67%
|
|--------------------------------------------------- | 73%
|
|-------------------------------------------------------- | 80%
|
|------------------------------------------------------------- | 87%
|
|----------------------------------------------------------------- | 93%
|
|----------------------------------------------------------------------| 100%
#> Warning: package ‘future’ was built under R version 4.5.2
#> Total computation time: 5.1 seconds (~ 0.08 minutes).
# Plot the membership stability
plot(fit_b, what = "stability", cutoff = 0.7)
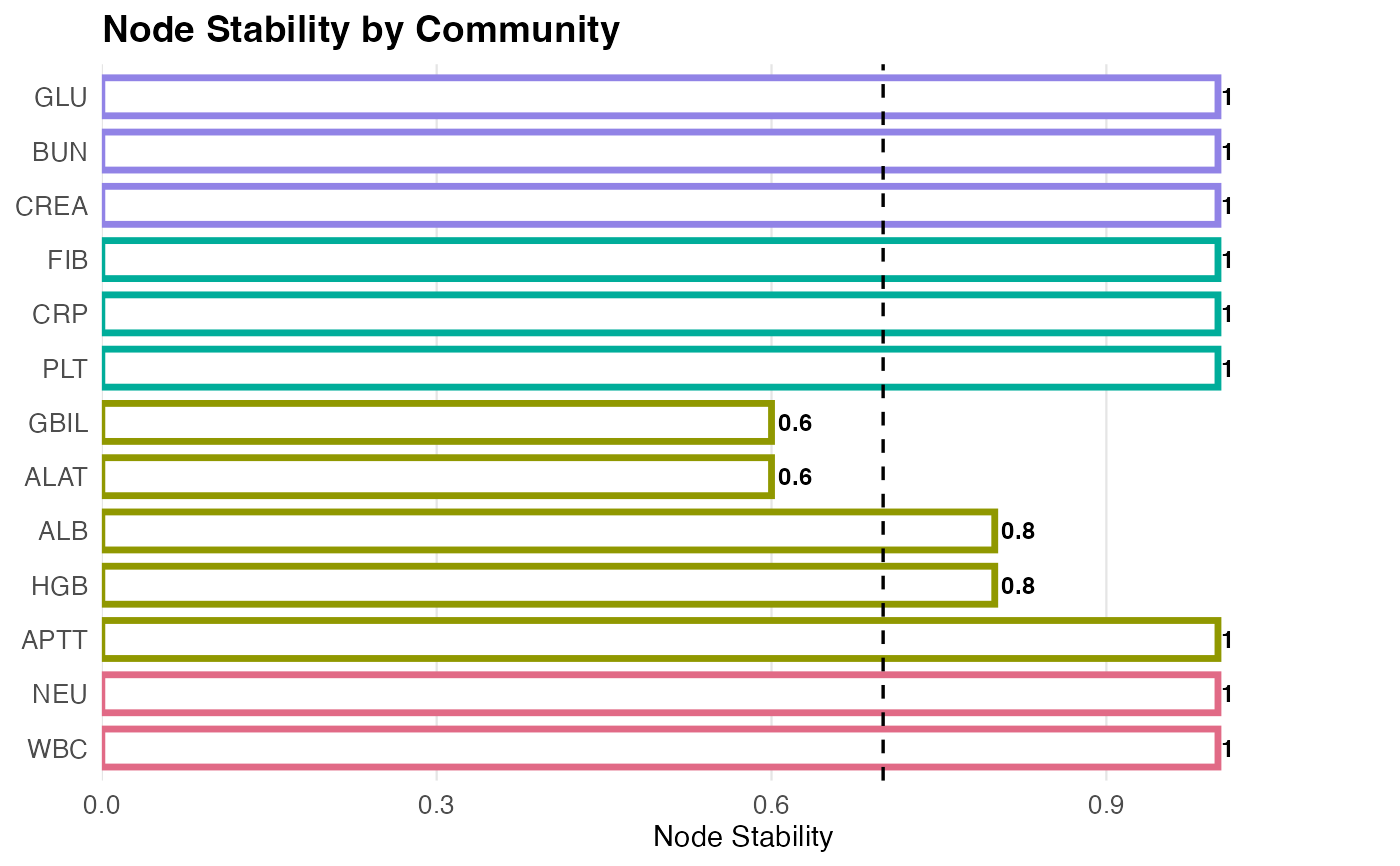 # }
# }